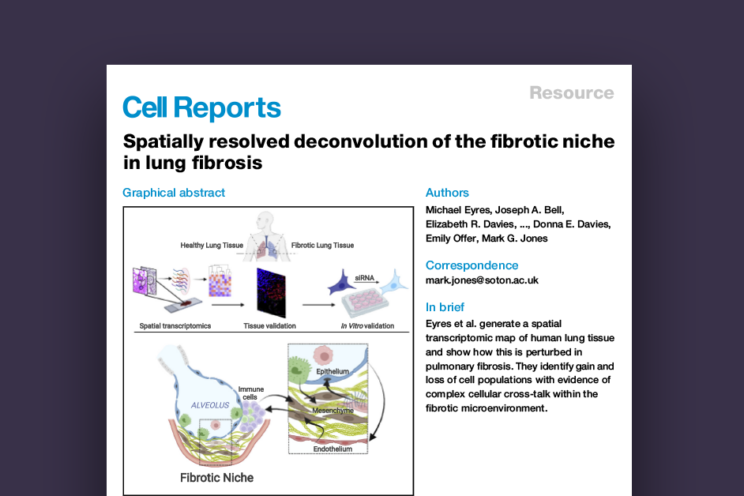
Spatially resolved deconvolution of the fibrotic niche in lung fibrosis
SUMMARY
A defining pathological feature of human lung fibrosis is localized tissue heterogeneity, which challenges the interpretation of transcriptomic studies that typically lose spatial information.
Here we investigate spatial gene expression in diagnostic tissue using digital profiling technology. We identify distinct, region-specific gene expression signatures as well as shared gene signatures. By integration with single-cell data, we spatially map the cellular composition within and distant from the fibrotic niche, demonstrating discrete changes in homeostatic and pathologic cell populations even in morphologically preserved lung, while through ligand-receptor analysis, we investigate cellular cross-talk within the fibrotic niche. We confirm findings through bioinformatic, tissue, and in vitro analyses, identifying that loss of NFKB inhibitor zeta in alveolar epithelial cells dysregulates the TGFb/IL-6 signaling axis, which may impair homeostatic responses to environmental stress. Thus, spatially resolved deconvolution advances understanding of cell composition and microenvironment in human lung fibrogenesis.