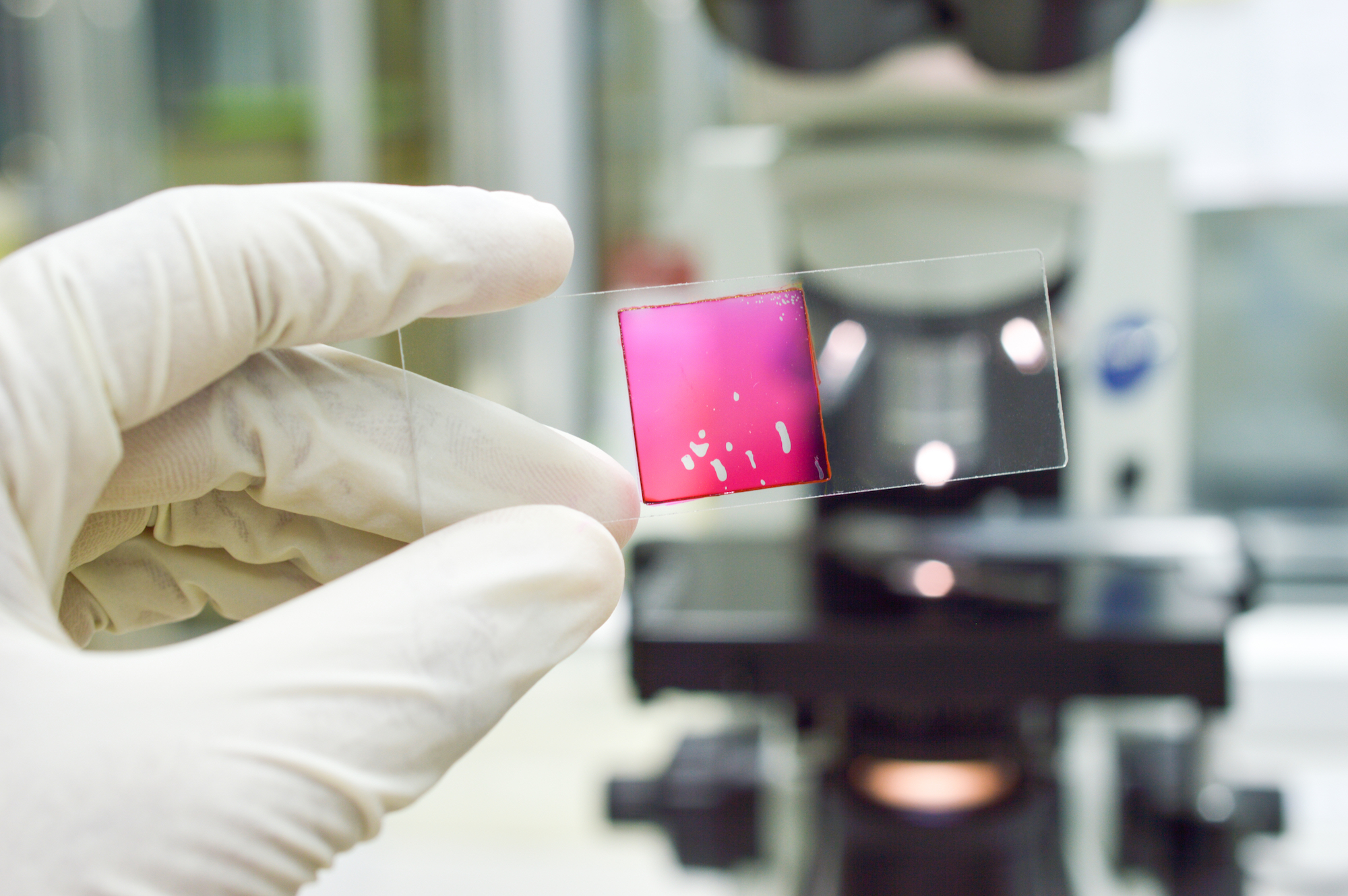
One of our collaborators, Intellegens, recently published a great paper in JCIM – Imputation of Assay Bioactivity Data Using Data Learning – focusing on applying Deep Learning approaches to tackle the problem of sparse data, a recurring issue in drug discovery.
Sparse data typically arise when compounds are being assayed against multiple biological targets, but not all compounds are assayed against all targets. Or, it can result from aggregating different literature studies with only partly overlapping compound sets and biological targets. A small sparse array of pIC50s might look like this:

The question then arises, can we make inferences about the missing data using the known data?
Can we predict, with any confidence, the missing values in this matrix based on ones we know? This would, for example, enable a project to reduce screening costs, or develop criteria to decide when to stop further screening or enable integration of multiple literature studies. This may appear to be a something-for-nothing aim but, the underlying latent structure in the data means that we can make surprisingly good estimates. But first, a detour to Netflix.
Look again at a data bioactivity array (Figure 2, below). Now replace the rows (proteins) with Netflix subscribers and replace the columns (compounds) with film titles, to generate Figure 3. Let each value in the array be a preference from 1 to 10, where 1 is hating the film and 10 is loving it.

This is the kind of data Netflix gets from user reviews – everyone has seen some films and reviewed them, but most individuals have not seen all films, and it would be very useful to be able to predict the missing preferences.
The potential for drug discovery
While the entertainment industry is a very different domain from drug discovery, the underlying problem here is the same – filling in the missing values – and the potential for borrowing methods used by them is there. Netflix were very keen to impute the missing values so that they could predict which film a subscriber would enjoy, recommend it to them and get another sale. It is the basis of the “if you liked that, you’ll like this” system.
We are just as keen to predict biological activities: “if your kinase liked that compound, it will love this one”
The success of this approach relies on underlying correlations between preferences of different users. Jim and Christine, from figure 3, both have similar film preferences. Therefore, it is likely Jim will like films that he has not seen, but that Christine has liked; in this case, Love Actually. Netflix use a technique based on matrix factorisation to extract this latent structure and make predictions.
Success with bioactivity data
Similar approaches have been used for bioactivity data with considerable success. If kinases A and B show similar sensitivity profiles, to compounds they have both been tested on, it is more likely that they will each be sensitive to compounds only the other has been tested against.
This factorisation method can only capture linear relationships between preferences, or data in general. Many problems, such as activity of a compound against a panel of biological targets, do not vary smoothly and generally display non-linear relationships.
Deep Learning is well-suited to dealing with non-linear relationships, let’s see what happened when Intellegens applied it to this bioactivity imputation problem.
Initially, they set the missing values to the mean of the known values; then iteratively improved the estimates using a deep neural network until convergence. They tested the method on a kinase data set: with 54% of the data used for model training, 14% for optimising hyperparameters and the remaining 32% as the cross validation set to test the model predictions in an unbiased manner.
Their approach outperformed matrix factorisation and traditional machine learning approaches, such as Random Forest, by a large margin. Performing similarly to a method developed at Novartis when predicting all the missing values.
A particularly attractive feature of the Intellegens approach is that, not only is the prediction made but, unlike the Novartis method, the confidence associated with each prediction can be generated. As a result, they could identify which predicted values were most likely to be correct. They were able to show that the R2, with the experimental values, rose from 0.445, when predicting all the values; through 0.7, when using the most confident 50% of predictions; to 0.9, when using only the most confident 1%. This allows screening resources to be focused on the most interesting or most uncertain values.
Seeing the applicability of their approach, more generally, whilst building a picture of how it behaves will be intriguing. Correlation Coefficients are a global estimate of model quality and the local behaviours around e.g. activity cliffs will be of interest.
About the author
Dr Andrew Pannifer is Lead Scientist in Cheminformatics at Medicines Discovery Catapult.
After a PhD in Molecular Biophysics at Oxford University, mapping the reaction mechanism of protein tyrosine phosphatases, he entered the pharmaceutical industry in 2002. Firstly at AstraZeneca and then at Pfizer, he performed structure-based drug design and crystallography, and in 2010 joined the CRUK Beatson Institute Drug Discovery Programme to start up Structural Biology and Computational Chemistry.
In 2013 he moved to the European Lead Factory as the Head of Medicinal Technologies to start up cheminformatics and modelling and also to work with external IT solutions providers to build the ELF’s Honest Data Broker system for triaging HTS output.