Why Electron Microscopy?
88% of the 210 drugs granted FDA approval between 2010 and 2016 were informed by structural biology. Traditionally structural biology has been underpinned by X-ray crystallography and to a lesser extent NMR. However, significant improvements in the field of Electron Microscopy (EM) has provided a powerful approach to studying protein structure.
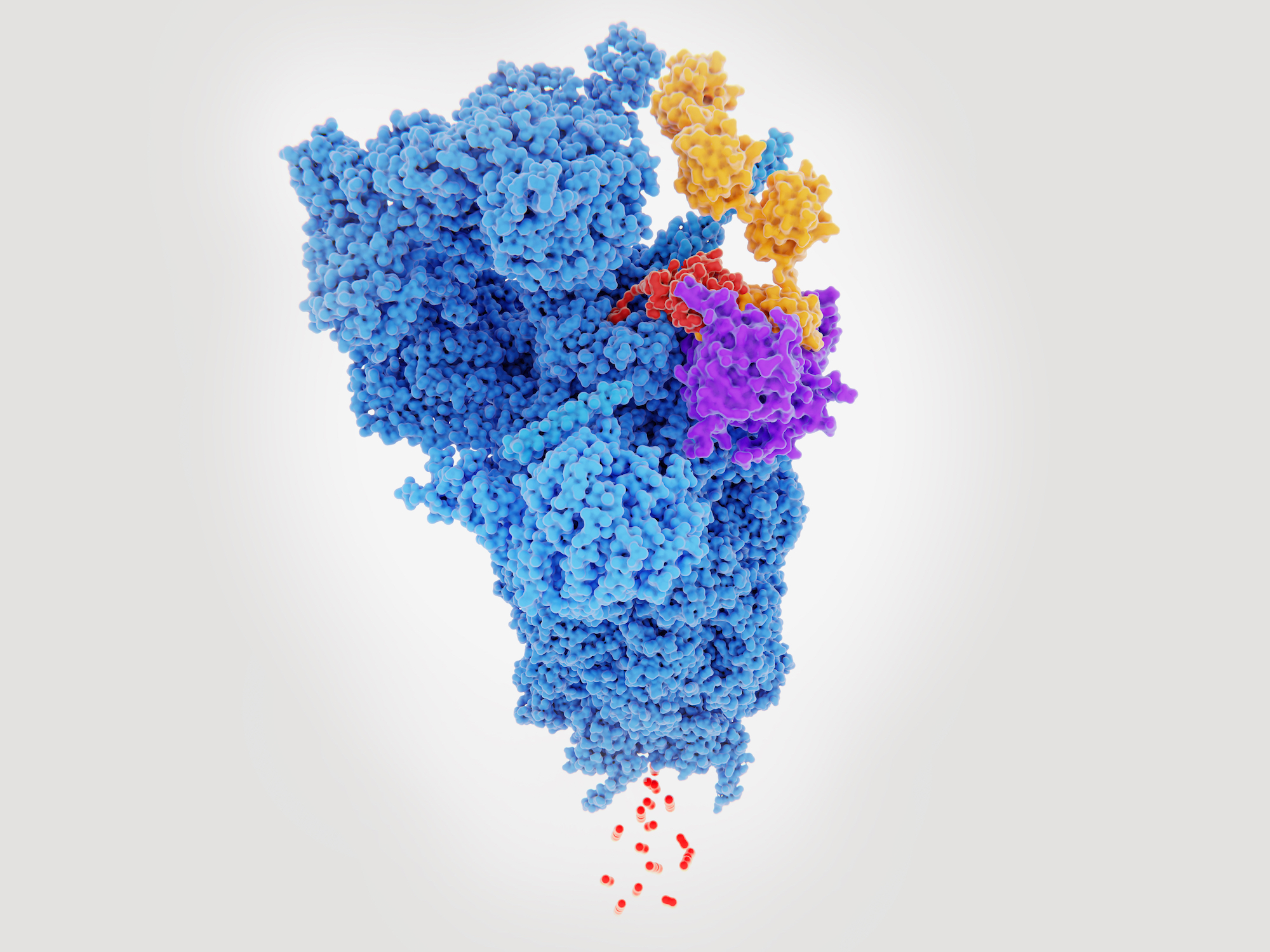
Traditionally EM was a low resolution technique that could only be used to study large protein systems >500kDa, but with recent developments resolutions as high as 1.6Å have been achieved and proteins as small as 50kDa have had their structures determined. This opens up a broad range of systems that can be studied spanning many clades of therapeutically important targets.
Changing the way we study membrane proteins
One of the most challenging protein targets are membrane proteins, which are often hard to express and stabilise outside of the membrane environment.
The traditional route of studying membrane proteins has been by X-ray crystallography, but this can rely on significant modifications to the proteins to make them amenable to crystallisation and is a very challenging process.
Despite ~60% of therapeutic targets residing in the membrane, they are poorly represented in terms of structures solved. In recent years, with the development in EM, there has been a rapid expansion in the membrane protein field with structures of key targets now available such as different members of the TRP family, hERG, LAT1, gamma secretase, Slo channels and the Ptch family. These new structures are underpinning therapeutic design and major pharma are now investing in their own EM facilities to complement existing structural biology groups.
More native like environments
Structural biology has traditionally been a reductionist discipline, relying on the over-expression of a protein, often lacking post translational modifications such as glycosylation. Proteins are then usually studied in an environment that best suits crystal growth or spectroscopic study (pH, temperature etc.). Moreover, these structures are often studied in the absence of binding partners and in the case of membrane proteins the native lipid environment.
To better understand the structure of a therapeutic target in the context of its native environment tomography can study proteins in situ. Although more limited in resolution this can provide a powerful approach to investigate not just the structure of a therapeutic target but also its wider environment that can have implications on structure and function. Moreover, tomography is a powerful approach for studying the wider effects of virus entry into cells or receptor signalling, for example.
Better understanding of different conformational states
In addition to improvements in the resolution from EM and the size of protein that can be studied, another powerful feature is the ability to pull out different conformational states. This has provided unprecedented detail into ion channels with the ability to capture open and closed states. This has important implications for structure based drug design approaches where different states may predominate in disease states, caused by a change in environment and / or mutations.
About the author
Dr Stephen Muench is a lecturer in membrane biology at the University of Leeds. His group primarily us Electron Microscopy (EM) to study the structure and mechanism of protein complexes, with a special interest in membrane proteins. His group work on a range of systems from ion channels to developing new biosensors.
He has been working on new ways to stabilise and study membrane proteins including the use of styrene maleic acid (SMA) co polymers to extract proteins with their closely associated lipids. Complementary to this his group are also developing new technologies to rapidly mix and freeze EM grids to trap different conformational states in the low ms time-frame (>10ms). This allows us to trap states that are poorly represented at equilibrium but may be important conformations for therapeutic intervention, such as probing different states of ion channel cycling.