PROTAC (PROteolysis TArgeting Chimera) is one of the hottest topics in drug discovery today as it is a highly promising new approach to targeted therapy. It offers several advantages over traditional occupancy-based drugs, with a potential to target the ‘undruggable’ proteome. Read our blog below to learn more about PROTAC and some assays that can be employed to measure protein degradation.
What is PROTAC?
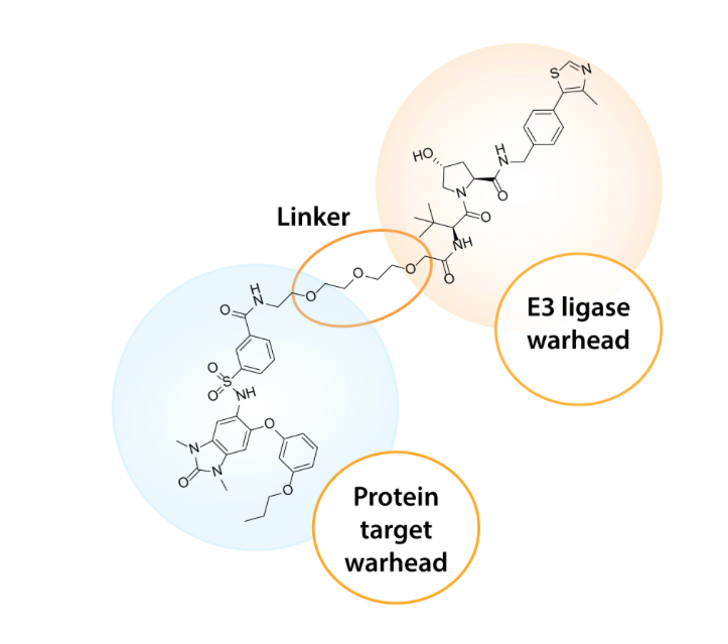
PROTAC is the ability to specifically degrade proteins using the endogenous machinery of the cell. The PROTAC molecule is designed with two ‘warheads’- one of them selectively binds to the disease target protein and the other to a specific E3 ligase to form a ternary complex. The two warheads are joined by a flexible PEG (Polyethylene Glycol) linker that brings them into close proximity. The E3 ligase recruits a ubiquitin loaded E2 conjugating enzyme to the ternary system, which then ubiquitinates the protein of interest. E2 enzymes keep getting added to the complex, resulting in poly- ubiquitination of the target protein. The poly-ubiquitinated protein gets recognised by 26S proteasome and subsequently gets destroyed by proteolysis. This approach involves sub-stoichiometric catalysis i.e. one degrader destroys many copies of the target protein. Once the protein is dispatched to the proteasome, the PROTAC molecule (degrader drug) dissociates from the complex, attaches itself to a new target and E3 ligase, and initiates another catalytic cycle. This process is known as ‘degrader recycling’.
PROTAC in Drug Discovery
PROTACs are highly appealing to drug developers as they have the potential to target the ‘undruggable’ proteome. Typically, the druggability of a protein depends on the inhibition of its activity. This is achieved by designing small molecules that can bind to a cavity or pocket in the protein of interest and block its function. However, there are several protein–protein interactions (PPIs) with flat protein surfaces and without deep pockets or well-defined binding sites, making them undruggable. PROTACs, on the other hand, regulate protein function by degrading target proteins instead of inhibiting them. They show better selectivity compared to inhibitors and avoid misinterpretations that arise from potential genetic compensation and/or spontaneous mutations owing to other gene editing tools. They are well distributed in the body and can be manufactured using suitable processes. Although they have relatively large molecular mass, PROTACs can endure sufficient intracellular concentrations of endogenous proteins, which helps them in protein degradation.
PROTACs show potential therapeutic benefits across a range of diseases such as cancer, viral infections, immunological disorders and neurodegenerative diseases. They can be administered orally, via injections or infusion. Arvinas, the pioneer of targeted protein degradation technology, demonstrated that PROTAC protein degraders can successfully penetrate the blood-brain barrier, which is vital for developing neurodegenerative drugs.
Despite the remarkable potential PROTACs have exhibited, challenges such as insufficient understanding of degradation activity and selectivity, limited structural data and lack of optimum computational tools still need to be tackled. It is therefore important that research continues in this field so the world can soon benefit from this new therapeutic category of drugs.
PROTAC assays
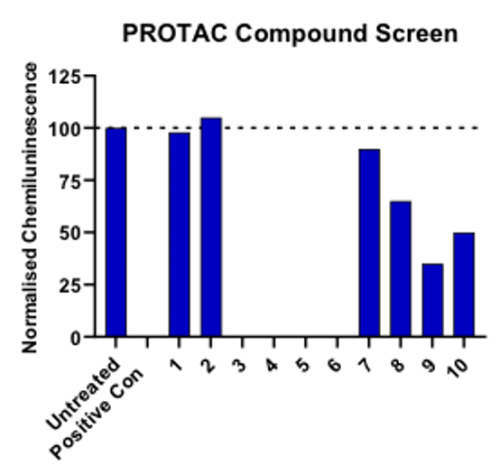
With all the attention PROTAC has been receiving lately, there is a strong need for robust assays to support the research in this area. Aurelia Bioscience, a UK based CRO, has successfully employed some screening strategies and methodologies for measuring protein degradation accurately:
1. WES and JESS
WES and JESS (ProteinSimple) are used to monitor target protein levels in cells following treatment with PROTAC test compounds. They have several advantages over traditional western blotting:
- Fully automated
- Highly reproducible
- Can run up to 25 samples plus a size ladder in just three hours
- Minimal sample volume is required: 3 to 5ul of 0.2ug/ul or above
- Can separate and analyse proteins by size from 2-440 kDa
Key factors for successfully quantifying protein degradation using WES and JESS include:
- A cell line with target expression at a relatively low level
- A high quality primary antibody
- A loading control that is not overly-expressed and can be multiplexed with the target antibody
- A well-defined methodology for PROTAC drug treatment and sample preparation
2. NanoLuc or HiBiT
Proteins can be labelled with either Nanoluc or HiBiT (Promega) and luminescence can be used to monitor protein levels in living cells. Following are some advantages of this technique:
- Live-cell kinetic studies of targeted protein degradation
- Quantitation of key degradation parameters
- Determination of kinetic profiles critical for understanding time frame of action
- Used with CRISPR to study endogenous proteins
- Excellent for phenotypic analysis
- Study of dynamic formation of ternary complex and protein ubiquitination
- Analysis of degradation rate and efficacy
- Assessment of PROTAC cellular permeability
Are you looking for a CRO to develop assays for your PROTAC project? Aurelia Bioscience has conducted studies with many PROTAC target molecules and can help you with your research. Get in touch with them to find out more >
About the author
Payash Bahuguna is Marketing Manager for Aurelia Bisocience.
She has a masters degree in Biotechnology and Entrepreneurship from the University of Nottingham and experience working with SMEs in the life science sector. Payash routinely writes blogs and articles for Aurelia Bioscience on both technical and non-technical subjects.